Reproducibility is an important aspect of research. One particular concept of interest is FAIR principles. FAIR stands for Findable, Accessible, Interoperable and Reusable and defines a ‘best practices’ approach for research.
However, complying with such a concept presents its own unique challenges: Some research tools rely on complex infrastructure elements and software stacks that are hard to replicate. In cases such as these, the added workload acts as an inhibitor on ensuring reproducibility. The result is that many experiments, data pipelines and results are hard to replicate. According to our preliminary insights, based on interviews within the ZHAW, on how experiments are conducted, such complexity is indeed commonplace in our current research activities.
It would thus be great, if the entire complex environment could be set up within a portable container, that anyone can simply download and use with no additional steps. Enter Docker. By no means ‘new’ technology, it has been around for several years and has been proven in many fields. Its well-documented advantages in software portability can also be leveraged to assist in creating portable and reproducible research software. A Dockerized research tool can run on any host that meets the hardware requirements with minimal setup, thus enabling a much simpler process of replicating the experiments and results of the original authors. There are already examples of leveraging Docker to improve reproducibility in research software, both internationally and within Switzerland. That said, eliminating the software setup is only one part of the equation.
In addition to making the software itself portable, the entire workflow of using it can be made reproducible with novel tools that enhance reproducibility. Examples include OSF and SDSC’s Renku. Renku is a tool designed entirely with the FAIR principles in mind, that augments the versioning functionality of git with the added benefits of a knowledge graph and other key features. Researchers can set up their repository much like with traditional git, with the added advantage of being able to record their entire workflow as they run it. This is stored in the repository’s knowledge graph and can be rerun on demand when new data or code changes are committed.
Both of these elements can be argued to provide an approach for increased reproducibility of research in isolation, nevertheless, we believe there are key ways in which they can complement each other to combat each other’s limitations.
Docker can make even some of the more complex tools reproducible, but researchers still need to learn how to operate the tool correctly to replicate a result.
Renku can record such a workflow and ‘replay’ it on command, assigning a much easier role to the human operator, but is restricted in the complexity of tools that it can natively manage.
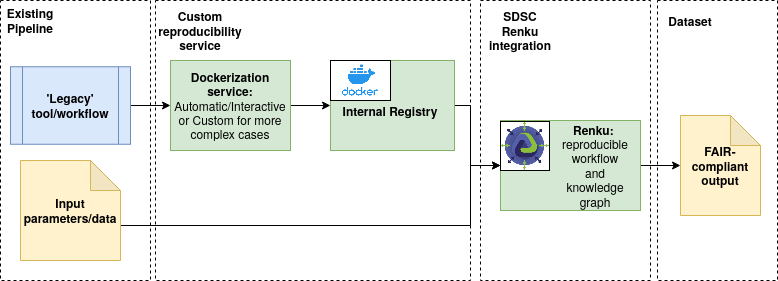
We thus see the benefit in a combined approach, with the development of customized ‘Renkurized’ images that contain the necessary software stack as part of the Docker image, while also giving access to the reproducible workflow functionality and knowledge graph of Renku. Such images would also become available in a dedicated catalogue, thus making the tools easily findable and accessible to all researchers and students. Public catalogues, such as Docker Hub are also viable and indeed there are interesting tools available there already, but we argue that to guarantee long-term availability to researchers even if the public portal were to move or change, a solution dedicated to our research is still the better choice.
Questions to be answered include how feasible it is to transform complex research setups into these custom container images, and whether this solution can be sustainable, with researchers within the ZHAW willing to shift to this approach for reproducibility in the future. We expect to answer these questions with an extended study of individual use-cases within the ZHAW.